Classification, names & identification
Classification, names & identification
Humans have a penchant for classifying living and non-living things. Things that are similar in some way can be grouped and given a name that allows them to be referred to by groups. Thus an art historian may talk of Impressionist painters, a geologist of sedimentary rocks and a zoologist of herbivorous animals. Impressionist, sedimentary and herbivorous are elements of classification schemes that are useful for particular purposes.
There are many ways of defining similarity for fungi. This leads to many useful ways of grouping them, with the groups created by one definition of similarity different to those created by another. An ecologist could find it useful to divide the fungi into those that are mycorrhizal and those that are not and each of those broad groups could be divided further. For example, while the mycelia of all mycorrhizal fungi form physical connections with plant roots, there are several different types of physical connections and the non-mycorrhizal fungi can be divided into sub-groups depending on how the mycelia feed. To an organic chemist a natural way to group fungi would be by the organic compounds they produce. Amatoxins (deadly to humans), anthraquinones (responsible for some of the bright pigments found in fungi) and lignin peroxidases (that decompose the complex lignin polymers in wood) are just three of the many classes of organic compounds found in fungi. These ecological and chemical groupings are very useful for certain purposes but they don't coincide. For example, you can find amatoxins in some mycorrhizal fungi as well as in some non-mycorrhizal fungi.
Nevertheless, when it comes to classifying living organisms many people are sure it has something to do with concepts such as species and genus and that it involves Latin names. The rest of this page will be devoted to the classification scheme built around concepts such these. There will be a look at some aspects of fungal classification and identification so that you can get an idea of how mycologists have gone about studying fungi over the past two centuries and there will be a few comments about the naming of fungi. However, there won't be a detailed discussion of the fine points of fungal classification - such as how different species or genera are defined. Nor will there be identification instructions.
|
Before going further it is worth pointing out the difference between classification and identification.
Classification answers questions of the sort: How is this fungus related to other fungi?
Identification answers the question: What's the name of the specimen in front of me?
Species, genus, the rest of the hierarchy - and naming
A common classification scheme for living organisms uses concepts such as species and genus (plural: genera) and in this scheme species is the basic unit. A genus typically contains a number of species and species is said to be of lower taxonomic rank than genus. Other examples of taxonomic rank in this classification scheme are family, order, class, phylum and kingdom - with each of higher rank than the one to its left (and family is of higher rank than genus). Hence there are fewer genera than there are species, fewer families than there are genera and so on. It is clear that this is a hierarchical scheme and any hierarchical system of classification is an example of a taxonomic classification. Any particular species, genus, family and so on is an example of a taxon (plural: taxa) . Thus Amanita muscaria ![]() , Amanita xanthocephala
, Amanita xanthocephala ![]() , Gymnopilus norfolkensis
, Gymnopilus norfolkensis ![]() and Hypholoma tuberosum
and Hypholoma tuberosum ![]() are examples of taxa at species rank. Amanita, Gymnopilus and Hypholoma are examples of taxa at genus rank. Amanitaceae and Strophariaceae are examples of taxa at family rank. The family Strophariaceae includes a number of genera including Stropharia, Hypholoma and Gymnopilus.
are examples of taxa at species rank. Amanita, Gymnopilus and Hypholoma are examples of taxa at genus rank. Amanitaceae and Strophariaceae are examples of taxa at family rank. The family Strophariaceae includes a number of genera including Stropharia, Hypholoma and Gymnopilus.
Various finer ranks are used at times. For example sub-species (which ranks below species) and sub-class (intermediate between order and class). However, while they are relevant to a specialist, they will be ignored here since they merely introduce technical complexities but illustrate no additional important principles.
A species' name consists of two words (genus name followed by species epithet) whereas the names of genera, families and so on are single words. There are rules governing the naming of fungal taxa (e.g. family names must end in 'aceae') and these rules are known as the International Code for the Nomenclature of algae, fungi and plants (or ICN), known before 2011 as the International Code of Botanical Nomenclature (or ICBN). Another ICN rule is that each taxon name must be attributed to its author (or authors), namely the person (or persons) responsible for the original description of the taxon. In 1987 Scott Redhead and Michael Kroeger published a joint paper in which they described the new species Hypholoma tuberosum so, in full, the name would be cited as Hypholoma tuberosum Redhead & Kroeger, the authors' names following the Latin species name. A little later there will be an additional note about what happens when a taxon name changes but otherwise this website omits the authors of taxon names since they are of relevance only to specialists. However, it is worth mentioning author names since you are likely to come across them when you look at other websites or through printed references. You can find the full name and author details for fungal taxa on the Mycobank website (http://www.mycobank.org/) .
.
Taxon names are treated as Latin words and so the rules of Latin grammar apply. Why Latin? The reason is largely historic, for Latin was the language of science in the 1700s and into the 1800s when the current system of naming came into existence and a great many taxon names were devised in those years. By the time Latin lost its role as the common scientific language it would have been an immense task to change every name - assuming there could be agreement as to which language to use in place of Latin. Though Latin is a dead language new words are often introduced from various non-Latin languages, 'Latinized' and then used. Latin taxon names are used for all groups of living organisms. For example, the native Australian animal pictured here ![]() is well-known as the Common Wombat and the word 'wombat' comes from an Australian aboriginal language. The scientific name of this species is Vombatus ursinus, the genus name being the Latinized version of 'wombat' and the species epithet is derived from a true Latin word meaning 'bear'.
is well-known as the Common Wombat and the word 'wombat' comes from an Australian aboriginal language. The scientific name of this species is Vombatus ursinus, the genus name being the Latinized version of 'wombat' and the species epithet is derived from a true Latin word meaning 'bear'.
Thus far we have a set of classification ranks and rules governing the naming of taxa at the different ranks. However the ICN is concerned only with the naming of taxa, not with how organisms are to be grouped into separate taxa. The creation of those rules required no knowledge about the organisms themselves. By contrast it's clear that separation into groups (i.e., deciding which organisms belong to which taxa) is possible only if you know something about the organisms' features. The ICN simply says that once you have created a hierarchical set of groups, by whatever means you choose, the ICN is the set of rules you will use for naming your groups. Two people may disagree strongly on how to create groups but both can follow the same rules when devising names for their separate collections of groups.
Evolution and grouping
Thoughts about how to group fungi into species, genera and so on have changed over time, though always with some idea of similarity involved but the earlier history of those changing ideas is irrelevant here. The later 1800s saw the beginnings of modern evolutionary theory with the core idea of descent with modification. Organisms pass on traits to their descendents but over time inherited traits may change and so organisms alive today may possess features markedly different from those of their ancestors many generations ago. If we knew how inherited traits had changed over time we could use such historical knowledge to produce a classification of organisms that would be consistent with evolutionary theory and from the later 1800s the goal has been to produce such a classification.
Let's illustrate this with a hypothetical example. Suppose that white, pink, green, rust brown, dark brown and black are the only spore colours found in mushrooms today. Suppose we know that all mushrooms had white spores until ten million years ago when green spores appeared in one population and all the descendents of this green-spored population have been green-spored. Some time later a population of the white spored mushrooms then extant gave rise to mushrooms with black spores. Later still some of the descendents of these mushrooms with black spores developed dark brown spores, while the other descendents continued to have black spores. Then some of the mushrooms with dark brown spores gave rise to mushrooms with rust brown spores while the other descendants had dark brown spores. Finally suppose that more recently still some of mushrooms still with white spores gave rise to mushrooms with pink spores and the descendents remained pink-spored. We could present this hypothetical scenario by the following diagram, where the arrow on the left indicates time.
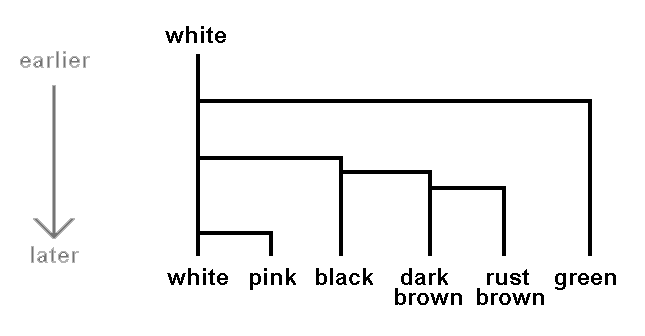
If, in this hypothetical scenario, today's mushrooms were divided into six basic groups based on spore colour it's clear that such a division would be consistent with evolutionary history. Another mycologist could argue that the shades of brown are insignificant and the brown-spored mushrooms should be considered as just the one basic group. There would then be five, not six, basic groups but both views are consistent with evolutionary history. Going back to six basic groups, two examples of higher level groups (or higher level taxonomic ranks) would be (rust brown, dark brown) and (rust brown, dark brown, black). Both are consistent with evolutionary history for the first of these arose from one offshoot from black-spored ancestors and the other from one offshoot from white-spored ancestors.
In the hypothetical example we had the benefit of perfect knowledge about the evolutionary history of spore colour but in real life perfect knowledge is never the case. In many other types of organisms there are fossils that can be used to help devise an evolutionary history but the fungal fossil evidence is very limited. Hence, if an evolutionary classification is to be produced, evolutionary history needs to be inferred, somehow, largely from what is found in fungi alive today. Fungi possess a variety of features that have been used to develop classification schemes and a number of these features will be presented a little later. For an evolutionary classification scheme it is necessary to know which form of each feature is the original form and which the changed forms. Moreover, it is necessary to know original forms for all fungi and also for various subgroups. In the hypothetical example white is the original form when we consider all the mushrooms but within the subgroup of brown and black spored mushrooms, black is the original form.
Sometimes there has been strong evidence for what was the original form of a trait but at other times the evidence has been ambiguous, leading to considerable debate. Not surprisingly, in such circumstances it is possible to devise more than one classification scheme, each using evolutionary principles but based on different hypotheses.
Changes in species names
If it were desired that species' names reflect an evolutionary classification then species' names would need to change as ideas about evolutionary relationships changed and that is what happens. Today Amanita xanthocephala ![]() is one of many species in the genus Amanita and Mycena interrupta
is one of many species in the genus Amanita and Mycena interrupta ![]() is one of many species in the genus Mycena. All those organisms with species names beginning with Amanita are considered to be evolutionarily closer to each other than any is to an organism that belongs to a species that has a name beginning with Mycena. Naturally another valid statement would be produced if the previous sentence were changed by swapping the words Amanita and Mycena. Had you been alive in 1860 you would not have seen those species' names in use but instead Agaricus xanthocephalus and Agaricus interruptus respectively. The change in names is a result of a better understanding of the evolutionary relationships amongst mushroom-forming fungi and a desire to give similar names to evolutionarily close species. Agaricus xanthocephalus is called a synonym of Amanita xanthocephala and Agaricus interruptus a synonym of Mycena interrupta .
is one of many species in the genus Mycena. All those organisms with species names beginning with Amanita are considered to be evolutionarily closer to each other than any is to an organism that belongs to a species that has a name beginning with Mycena. Naturally another valid statement would be produced if the previous sentence were changed by swapping the words Amanita and Mycena. Had you been alive in 1860 you would not have seen those species' names in use but instead Agaricus xanthocephalus and Agaricus interruptus respectively. The change in names is a result of a better understanding of the evolutionary relationships amongst mushroom-forming fungi and a desire to give similar names to evolutionarily close species. Agaricus xanthocephalus is called a synonym of Amanita xanthocephala and Agaricus interruptus a synonym of Mycena interrupta .
It is a convenient memory aid if evolutionarily close species have the same genus name. However, division of organisms into groups and the naming of those groups are two different processes and there need be no connection between them. This is certainly the case with many folk names. Fly Agaric and Deathcap Mushroom are vernacular names, the first for those organisms given the species' name Amanita muscaria and the second for those given the species' name Amanita phalloides. The two vernacular names are well-established and unambiguous but give no indication of the nature of the relationship between those organisms called Fly Agarics and those called Deathcap Mushrooms. Incidentally, at one time the species' names Agaricus muscarius and Agaricus phalloides were in use and in principle there is no reason why they could not be used today just as names, but no longer with any classificatory significance in the Agaricus part of the name.
|
Earlier I noted that the ICN rules require any taxon name be attributed to its author (or authors) but at that stage made no mention of what happens when there is a change in taxon name. The species Agaricus interruptus was described by Miles Berkeley in 1859 and was re-classified to the genus Mycena in 1887 by Pier Andrea Saccardo. Today, you would see the full citation as Mycena interrupta (Berk.) Sacc., with the names of Berkeley and Saccardo present, but usually in abbreviated form as given here. Berkeley recognized that the small blue mushrooms represented a taxonomic group distinct from any of the then known taxonomic groups and gave the group the name Agaricus interruptus. Saccardo also thought that the small blue mushrooms represented a taxonomic group distinct from any other group but in his view they were better classified in the genus Mycena. The change in species epithet from interruptus to interrupta is simply a requirement of Latin grammar. There had been no change in the way that the taxonomic group was defined, simply a change in the name given to the group. The name Berkeley has been kept (in brackets) to acknowledge that he was the first to define this taxonomic group. Here's another example, involving the luminous fungus pictured below. It has been known by the names Agaricus nidiformis Berk., Pleurotus nidfiformis (Berk.) Sacc. and Omphalotus nidiformis (Berk.) O.K. Mill., the name by which it is currently known. As you'd surmise the original description was by Berkeley (in 1844). There was a re-classification by Saccardo (in 1887) and another re-classification by Orson Miller in 1994. Once again there have been no changes in the concept of the taxonomic group, just in the name. As in the Mycena example, Berkeley is kept (in brackets) and is followed by the name of whoever is responsible for the latest re-classification.
Features that have been used in fungal classification
At any time mycologists can use only the tools available to them and, not surprisingly, this means that the features used in classification have changed over time as more has been discovered about fungal structure or biology. Features that are thought significant at one time may later be found to be unreliable. In this long section I will mention some of the tools and features that have been used to generate fungal classification schemes.
The early classification schemes relied heavily on MACROSCOPIC FEATURES of the fruiting bodies. However, certainly by the late 1800s it had become clear that fruiting bodies could sometimes show variation in form, depending on external conditions, and clearly it is pointless to base a classification on any features subject to change by external conditions. The following pair of photos shows an example of marked macroscopic variation. Both show the species Flammulina velutipes. The white form is sold commercially as enokitake in Asian grocery stores and the other photo is of the species growing in the wild. Apart from the colours the wild form has a stem with a velvety feel. Here ![]() are two illustrations of the species, showing the short bristles on the stems. To produce the enokitake form the mycelium is grown in the dark in a jar of sterilized sawdust. Around the mouth of the jar there is a long plastic collar that induces the formation of the elongated stems.
are two illustrations of the species, showing the short bristles on the stems. To produce the enokitake form the mycelium is grown in the dark in a jar of sterilized sawdust. Around the mouth of the jar there is a long plastic collar that induces the formation of the elongated stems.
 |
Both photos show Flammulina velutipes. The white version is the form sold commercially as enokitake. |
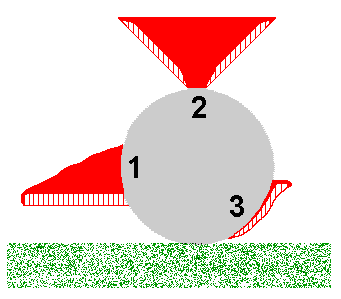 You may object to the enokitake example on the grounds that it's very artificial and so isn't highly relevant to what happens in the real world. In answer to that I note that aberrant forms of fruiting bodies of various fungal species have been found in naturally dark habitats such as mines or caves. But let's go out into the light and consider the polypores. Polypore fruiting bodies are generally corky to woody in texture and many polypore species usually produce shelf-like fruiting bodies, such as shown by the stylized cross-sectional figure 1 in the accompanying diagram. This figure represents a fruiting body growing out of a fallen log, shown in cross-section in grey. In figure 1 the fertile layer of basidium-bearing tubes is at the bottom of the fruiting body and the bulk of the 'shelf' is the solid red, sterile supporting tissue. For optimum spore release the tubes need to be vertical and the shelf-like fruiting body ensures the vertical orientation of the tubes. Suppose that for some reason the development of the fruiting body had been triggered at the top of the log. In that case, the need for the tubes to be vertical could induce a fruiting body of the type shown in figure 2. Finally, if the fruiting body development were triggered near the base of the log, we could get a fruiting body as shown in figure 3. A classification based on macroscopic features could lead to the fruiting bodies in figures 1, 2 and 3 being assigned to three different species, perhaps in different genera, even though all were produced from the one mycelium. Pycnoporus is a polypore genus and the orange fruiting bodies are usually shelf-like outgrowths from dead wood, with many shown here
You may object to the enokitake example on the grounds that it's very artificial and so isn't highly relevant to what happens in the real world. In answer to that I note that aberrant forms of fruiting bodies of various fungal species have been found in naturally dark habitats such as mines or caves. But let's go out into the light and consider the polypores. Polypore fruiting bodies are generally corky to woody in texture and many polypore species usually produce shelf-like fruiting bodies, such as shown by the stylized cross-sectional figure 1 in the accompanying diagram. This figure represents a fruiting body growing out of a fallen log, shown in cross-section in grey. In figure 1 the fertile layer of basidium-bearing tubes is at the bottom of the fruiting body and the bulk of the 'shelf' is the solid red, sterile supporting tissue. For optimum spore release the tubes need to be vertical and the shelf-like fruiting body ensures the vertical orientation of the tubes. Suppose that for some reason the development of the fruiting body had been triggered at the top of the log. In that case, the need for the tubes to be vertical could induce a fruiting body of the type shown in figure 2. Finally, if the fruiting body development were triggered near the base of the log, we could get a fruiting body as shown in figure 3. A classification based on macroscopic features could lead to the fruiting bodies in figures 1, 2 and 3 being assigned to three different species, perhaps in different genera, even though all were produced from the one mycelium. Pycnoporus is a polypore genus and the orange fruiting bodies are usually shelf-like outgrowths from dead wood, with many shown here ![]() . The upper surfaces of the older fruiting bodies shown there have become bleached. Here
. The upper surfaces of the older fruiting bodies shown there have become bleached. Here ![]() is a fruiting body that has grown out from the top of a fallen log.
is a fruiting body that has grown out from the top of a fallen log.
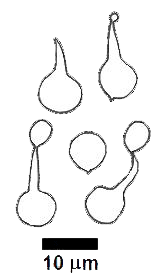
This led to a greater reliance on microscopic features in classification, since these were not modified by external factors. Of course, a feature that is of little value in classification can still be very useful in identification. There's no paradox in that statement and there's more about this aspect in the final section of this page. The nature of the asci and basidia were taken as fundamental features in the classification of the ascomycetes and basidiomycetes. For example, while the basidia in most basidiomycete species are single-celled and four-spored there are variations within just this form but there are also other forms of basidia and the following drawings show some of the variety.
There is also considerable variety in spores, for they may be smooth or ornamented, vary in shape from spherical to highly angular, be white or coloured when viewed in mass (as in a SPORE PRINT) and one-celled or septate. The following drawings show some of the variation of form in spores.
These drawings show (from left to right) the spores of Gloniopsis praelonga, Tubulicium vermiferum, Dacryobolus sudans, Hydnangium carneum and Calostoma fuscum. |
There are also other microscopic features that have been used and since polypores have been mentioned I'll note that one of the microscopic features that was used to help classify polypores was the nature of the hyphae. Early in the 20th century it was found that the fruiting bodies of some polypores were composed of only one type of hypha, some of two types and some of three. The hyphal construction of the fruiting bodies was not influenced by outside factors and the hyphal nature of the fruiting bodies could be used to help differentiate species in a reliable way so that hyphal analysis had become an important tool in polypore identification and classification by the mid-20th century.
These comments about the importance of microscopic features do not mean that macroscopic features were ignored, just that various macroscopic features declined in classificatory significance. Some lost all value while others were still used but for purposes different to before. I will look at some basidiomycete examples. Certainly by the beginning of the 20th century it had become clear to a number of mycologists that there could be close relationships between fungi of quite different fruiting body types. Consider the genera Tremella (fruiting body three dimensional, convoluted and gelatinous) and Exidiopsis (fruiting body flat, thin and waxy to dry). Both have the Tremella-type basidium, shown in the drawings above, and Narcisse-Théophile Patouillard (1854-1926), a pioneer in the microscopic study of basidiomycetes, classified them close together. Incidentally, not all genera with gelatinous-textured fruiting bodies have Tremella -like basidia. The earlier example of three polypore fruiting bodies growing from a fallen log indicated that fruiting body form need not be a foolproof method of distinguishing genera within the polypores but at least each of those fruiting bodies would have polypore features (e.g. a corky to woody texture). Thus, through much of the 20th century it still seemed sensible to think of the polypores as a natural grouping. Similarly most of the mushroom species seemed to constitute a natural grouping, though some exceptions were recognized. For example, the mushroom genera Paxillus and Phylloporus share a number of microscopic and macroscopic features with the boletes and some mycologists favoured classifying these genera with the mushrooms while others favoured putting them in with the boletes. The fleshy coral fungi and the puffballs were two more examples of what seemed natural groupings amongst the basidiomycetes.
|
In the 20th century the development of electron microscopes allowed more detailed studies of microscopic structures. For example spore walls in many species were shown to be composed of several distinct layers and there was variation in the complexity of the walls between neighbouring cells in a hypha. However, classification schemes were not restricted to the use of structural features, whether macroscopic or microscopic. From the later 1800s onward there were numerous laboratory studies of life cycles in which spores were germinated and the processes of mycelial growth carefully followed and fruiting body development carefully observed. In most species spores always germinate to produce a mycelium, as shown by the diagram near the beginning of the MYCELIUM page. A small number of basidiomycete genera can sometimes display a phenomenon known as germination by repetition in which a spore, instead of producing a mycelium, produces a short angular projection and atop that a secondary spore, of a form similar to the first. In effect the original spore behaves like a miniature, one-prong basidium, This phenomenon is found in several genera with three-dimensional, gelatinous fruiting bodies and septate basidia as well as in several corticioid genera (with flat, dry to waxy fruiting bodies) some with septate basidia and some with non-septate basidia. You can see that germination by repetition is a phenomenon that correlates neither with the macroscopic feature of fruiting body type nor with the microscopic feature of basidium type. It is an unusual phenomenon in fungi and clearly gives yet more food for thought as to how the various combinations of fruiting body type, basidium type and germination by repetition evolved - and hence how all that information can be used in classification.
Now suppose you have some mycelia growing in your laboratory and these produce fruiting bodies. In many cases these would be of the same sort from which you collected the spores for your experiments, but occasionally you may see aberrant forms and these can be informative. As an example I will mention some experiments involving Deconica merdaria (once known as Psilocybe merdaria), which produces small mushrooms. Mycelia that grew from spores gathered from a spore print developed several types of fruiting body. The three forms produced most consistently were (1) the standard mushroom form; (2) a mushroom form but with a lateral rather than central stem and (3) a stem-less and more-or-less globose fruiting body, somewhat truffle-like. Given what was said earlier about enokitake it is natural to wonder if the laboratory conditions were such that they alone brought about the aberrant forms, but normal forms were produced at the same time. The experimenter also grew mycelia from spores of several other mushroom species in the same laboratory conditions but these produced only the standard, mushroom-shaped fruiting bodies. This indicates that Deconica merdaria could produce mushrooms or somewhat truffle-like fruiting bodies under similar conditions and such a finding naturally prompts questions about the relationships between mushrooms and the basidiomycete truffle-like species. Laboratory experiments are useful in that you can produce numerous fruiting bodies at leisure and the more you produce the greater the chance of seeing something unusual. However, a lot of people have spent a lot of time looking at fungi in the field and a variety of aberrant forms of fruiting bodies of various species have been found in the wild, at times growing alongside normal forms .
.
As you can see from the examples of Dermocybe splendida ![]() and these species of Entoloma
and these species of Entoloma ![]() , Mycoacia
, Mycoacia ![]() and Phanerochaete
and Phanerochaete ![]() , a variety of pigments are found in fungi. These pigments are examples of the chemical compounds produced by fungi and if two fungi produce identical or very similar compounds it is natural to wonder if such chemical similarity indicates a close evolutionary relationship. Chemical analyses have been used in fungal classification, though most commonly in the study of lichens (also known as lichenized fungi) and there's more about this on the CHEMISTRY page of the ANBG lichen website (http://www.anbg.gov.au/lichen/). Chemistry has been a common tool at the micro level in that there are some standard chemical tests that are applied to spores, asci and various other microstructures. For example, if a fungus has colourless spores (when they are mounted in water on a slide and viewed through a microscope) it is standard procedure to mount some spores in Melzer's reagent (essentially an iodine solution) and see if anything happens. Melzer's reagent is a brownish yellow liquid and if the spores show no reaction they take up that yellowish brown colour. However, there are species in which the spores turn some shade of blue in Melzer's (called an amyloid reaction) and other species in which the spores turn some shade of red (a dextrinoid reaction). Melzer's reagent is used on micro-structures other than spores and it has been used both in classification and identification. This reagent is a major chemical tool but various other chemical reagents are also in use, some with a variety of roles others only for very specific tests. Some have interesting ingredients and an example is sulphovanillin - vanillin in concentrated sulphuric acid. Nasty, but it does have a very nice smell!
, a variety of pigments are found in fungi. These pigments are examples of the chemical compounds produced by fungi and if two fungi produce identical or very similar compounds it is natural to wonder if such chemical similarity indicates a close evolutionary relationship. Chemical analyses have been used in fungal classification, though most commonly in the study of lichens (also known as lichenized fungi) and there's more about this on the CHEMISTRY page of the ANBG lichen website (http://www.anbg.gov.au/lichen/). Chemistry has been a common tool at the micro level in that there are some standard chemical tests that are applied to spores, asci and various other microstructures. For example, if a fungus has colourless spores (when they are mounted in water on a slide and viewed through a microscope) it is standard procedure to mount some spores in Melzer's reagent (essentially an iodine solution) and see if anything happens. Melzer's reagent is a brownish yellow liquid and if the spores show no reaction they take up that yellowish brown colour. However, there are species in which the spores turn some shade of blue in Melzer's (called an amyloid reaction) and other species in which the spores turn some shade of red (a dextrinoid reaction). Melzer's reagent is used on micro-structures other than spores and it has been used both in classification and identification. This reagent is a major chemical tool but various other chemical reagents are also in use, some with a variety of roles others only for very specific tests. Some have interesting ingredients and an example is sulphovanillin - vanillin in concentrated sulphuric acid. Nasty, but it does have a very nice smell!
Suppose that species X is reported to occur in both Europe and Canada. It is natural to wonder if this is really a case of one disjunct species or are there in fact two very similar species, one on Canada and the other in Europe. Another type of question arises when species A and species B, very similar to each other, have overlapping distributions. Are they really two distinct species? Some notable examples of the A/B scenario have involved wood-inhabiting corticioid fungi in the Northern Hemisphere. The corticioid fruiting bodies are flat with little in the way of macroscopic features so identification relies on microscopic features. There have been pairs of corticioid species, the fruiting bodies macroscopically identical, with some microscopic differences but with the major difference being that one of the pair occurs only on coniferous wood and the other only on broadleaf wood. In such a case it is natural to wonder if there are really two species, with different substrate requirements, or the one species that can tolerate any woody substrate. Questions such as these have arisen often and over the past century mating studies have been carried out to help find the answers. The effectiveness of mating studies relies on the fact that in many species a mycelium that develops from a single spore will not produce fruiting bodies. Fruiting bodies will form only from a composite mycelium that develops once two single-spore mycelia have come into contact and the composite mycelium is microscopically (and often macroscopically) distinct from the single-spore mycelia. Let two spores germinate a little distance apart on an agar plate. The two single-spore mycelia will grow radially, eventually come into contact and if, after a while, you can detect a composite mycelium, you can be sure that the two spores came from the same species. Success in forming a composite is immediately informative whereas one failure to form is not. After all, perhaps a composite failed to form because of some technical problem rather than being the result of the spores coming from different species. In practice there would be replications of such experiments to guard against problems with the experimental set-up, but the simple outline given above explains the essence of a mating study. So, to answer the questions raised early in this paragraph, get spores of species X from both Canada and Europe and also spores of A and B and follow the process described above.
More recently there has been much emphasis on analysis of genomes. An organism's genome is its hereditary information, stored as DNA, and containing that organism's genes. The fact that DNA is inherited from parents and is subject to change over time, makes genomic analyses attractive as a means of determining genetic and hence evolutionary closeness. In such analyses one looks at equivalent sections from the genomes of representative specimens from different species and the results are presented as sequences of the letters A, C, G and T, indicating how the four basic DNA components or bases (adenine, cytosine, guanine, thymine) are arranged. If you have DNA sequences from representative specimens of different species you can look for the differences between them. For example, suppose that the following are corresponding sections from the sequences of three species:

In this section of DNA the three sequences are very similar, with the differences between sequence 1 and sequence 2 shown in red, and the differences between sequence 2 and sequence 3 shown in blue. You could say that, with regard to this section of DNA, species 1 is closer to species 2 than it is to species 3 since it takes three changes to transform sequence 1 into sequence 2, but five changes to transform it into sequence 3. This immediately suggests the possibility of defining a measure of distance between two sequences based on the number of differences in their bases. Then it would seem a natural extension to use the distances between sequences as proxies for measures of evolutionary closeness between species so, in the above example, species 1 would be evolutionarily closer to species 2 than to species 3. In fact there are various definitions of distances between sequences and while the simplistic scenario given here is sufficient to explain the basic idea there are complications and pitfalls to be aware of if a meaningful distance measure is to be produced. Another point to remember is that while a distance measure gives a useful summary statistic about a pair of sequences, it discards the information within each sequence.
Thus far everything has been based on the transmission of genetic material from parent to offspring, referred to as vertical gene transfer. There is another phenomenon, known as horizontal gene transfer (or lateral gene transfer). in which genetic material is gained from a non-parental organism, indeed perhaps from a markedly different organism. For example, it has been shown that production of carotenid pigments by pea aphids relies on genes that are most likely to have been gained by an aphid ancestor through horizontal transfer from fungi. It is natural to wonder how the existence of horizontal gene transfer influences ideas of evolutionary closeness. After all, similarity between the DNA sequences of two species could be the result of horizontal transfer from one to the other (or into both from a third) rather than an indicator of evolutionary closeness. Horizontal gene transfer is common amongst the prokaryotes (organisms without nuclei, such as bacteria), at times with the horizontal transfer of large amounts of genetic material. Eukaryotes are organisms with nuclei and fungi, plants and animals are examples. The evidence indicates that eukaryotes are far less commonly involved in horizontal gene transfer and when eukaryotes are the recipients (from either other eukaryotes or prokaryotes) the amount of transferred genetic material is small. Clearly it is essential to distinguish the effects of horizontal and vertical transfer but for the purpose of this page it is not necessary to go into the details of how that is done .
.
Tree diagrams
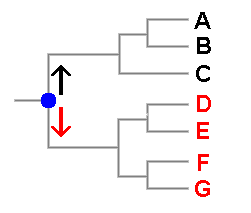 Tree diagrams are used frequently in books, papers or websites to display evolutionary relationships between species and there is a hypothetical example to the right of this paragraph in which six species, denoted A to G are shown. You can see that it is very similar to a human family tree. In fact if this were such a family tree you'd say that A and B were siblings, and hence more closely related to each other than either is to C. You'd also say that back in time at the point marked in blue two lines of descent arose, one leading to the three black people and the other to the four red people. Another way of looking at this is to say that the blue dot indicates the most recent ancestor common to the black and red people. In a similar vein it is possible to say that two species are more closely related to each other than either is to any other and it makes sense to talk of the most recent common ancestor of a group of current species. A tree such as this shows hypotheses about the species' evolutionary relationships, or phylogeny, but a phylogenetic tree is not itself a classification though it can suggest possible classifications. Here, for example, you could group the black species into one genus and the red species into a second. Someone else might think that three genera are warranted, with D,E separate from F,G. As in the earlier spore colour example, the two views given here are evolutionarily plausible. Phylogenetic trees can be produced from various types of data but now most trees are based on DNA sequence data and there are various methods of using sequence data to obtain trees. Some require the calculation of distances between sequences and use only the distance information to build up a tree. Other methods (involving the concepts of parsimony or likelihood) don't build a tree but choose a tree from all possible trees. For any group of objects there is only a finite number of ways in which those objects can be organized into trees. There are various ways of assigning a score to each possible tree and in principle it is then a matter of selecting, from all possible trees, the one with the best score. Of course, the phrase 'in principle' in the previous sentence hides a variety of potential complications but they are irrelevant to this page.
Tree diagrams are used frequently in books, papers or websites to display evolutionary relationships between species and there is a hypothetical example to the right of this paragraph in which six species, denoted A to G are shown. You can see that it is very similar to a human family tree. In fact if this were such a family tree you'd say that A and B were siblings, and hence more closely related to each other than either is to C. You'd also say that back in time at the point marked in blue two lines of descent arose, one leading to the three black people and the other to the four red people. Another way of looking at this is to say that the blue dot indicates the most recent ancestor common to the black and red people. In a similar vein it is possible to say that two species are more closely related to each other than either is to any other and it makes sense to talk of the most recent common ancestor of a group of current species. A tree such as this shows hypotheses about the species' evolutionary relationships, or phylogeny, but a phylogenetic tree is not itself a classification though it can suggest possible classifications. Here, for example, you could group the black species into one genus and the red species into a second. Someone else might think that three genera are warranted, with D,E separate from F,G. As in the earlier spore colour example, the two views given here are evolutionarily plausible. Phylogenetic trees can be produced from various types of data but now most trees are based on DNA sequence data and there are various methods of using sequence data to obtain trees. Some require the calculation of distances between sequences and use only the distance information to build up a tree. Other methods (involving the concepts of parsimony or likelihood) don't build a tree but choose a tree from all possible trees. For any group of objects there is only a finite number of ways in which those objects can be organized into trees. There are various ways of assigning a score to each possible tree and in principle it is then a matter of selecting, from all possible trees, the one with the best score. Of course, the phrase 'in principle' in the previous sentence hides a variety of potential complications but they are irrelevant to this page.
The Inkcaps
Often new evidence suggests a close relationship between species previously thought not to be closely related and there are some examples of this phenomenon on the RELATIONSHIPS THAT ARE AND ARE NOT page. The opposite also happens, with the new evidence indicating that species previously thought to be closely related are not so close and this section will deal with an example, the Inkcap mushrooms. There are a number of mushroom-producing fungi in which the mushroom caps dissolve into a black, inky mess. Over time, perhaps hours, perhaps days (depending on species and weather conditions) the cap of such a species disappears, starting from the outer edge, until only a small central portion is left atop the stem. Probably the best known of these is the Shaggy Inkcap (Coprinus comatus). Shown here ![]() is a group of Shaggy Inkcaps, still all white, before the inkiness has set in. The photo to the right of this paragraph shows one with half the cap gone. You can see the black edge to the cap and on the left a couple of inky blobs hanging from the cap edge and soon to fall off. In this species inkiness is associated with spore dispersal and there is a section about spore release in Inkcap mushrooms on this page.
is a group of Shaggy Inkcaps, still all white, before the inkiness has set in. The photo to the right of this paragraph shows one with half the cap gone. You can see the black edge to the cap and on the left a couple of inky blobs hanging from the cap edge and soon to fall off. In this species inkiness is associated with spore dispersal and there is a section about spore release in Inkcap mushrooms on this page.
Inkiness is both a very striking visual feature and one that is rare in fungi. Therefore it is not surprising that inkiness was taken to be an indicator of a close relationship and in earlier times the great majority of the species that dissolved were considered to belong to the genus Coprinus. Over time the genus Coprinus grew to include over 150 species, some with the inkiness slight to non-existent. However late in the 1900s the initial DNA studies indicated that the species in Coprinus were not related closely enough for all to be placed in the one genus - in fact, not even in the one family. This was also the conclusion even if only the definitely inking species of Coprinus were considered. For example, on the basis of the molecular evidence, Coprinus comatus was more closely related to some other, non-inking genera, than it was to many other inking species of Coprinus. These DNA findings were controversial and a number of mycologists thought the implications nonsensical, being contrary to long-held ideas . What do you do when you get new evidence that contradicts existing concepts? Obviously, re-check the methods to see if there have been any mistakes. If not, you can either accept one lot of evidence as more reliable than the other or leave the issue unresolved. Not necessarily a very happy result, but sometimes it's necessary to put a problem aside and wait for future developments to resolve the issue. In the case of Coprinus, there was further DNA analysis, using improved techniques and with analyses of larger segments of Coprinus genomes, and the conclusions were much the same. One thing to note is that the DNA evidence didn't come as a surprise to some mycologists, since there had been debate (for over a hundred years) about the correct relationships between the Coprinus species. Long before DNA analyses became possible a number of mycologists had pointed out that, based on various microscopic features, Coprinus was a mixed bag.
Don't think that the introduction of DNA sequence analyses into taxonomy was the first time a new method of investigation had generated controversy. Each new tool or method has provided information previously unobtainable and in each case the new evidence has supported some aspects of contemporary classification schemes but contradicted others and led to various mycologists strongly arguing against the usefulness of the newer techniques.
The DNA results prompted re-assessment of the macroscopic and microscopic structures of the Coprinus species to see if any structural features were correlated with the DNA evidence and that was found to be the case. The DNA evidence indicates that Coprinus comatus and a few other species form a closely related group, so there's a good argument for keeping these together. Apart from the DNA evidence, the species in this group share some microscopic and macroscopic features that were not found in other Coprinus species. One macroscopic feature is very easy to see. The stem of Coprinus comatus is pipelike, rather than solid, but the pipe isn't empty. There's a thin cord, composed of a bundle of hyphae, that runs the length of the hollow centre and has no known purpose. This photo ![]() shows a dried, herbarium specimen of Coprinus comatus, with the stem cut open to reveal the wispy central cord and there's a closer view here
shows a dried, herbarium specimen of Coprinus comatus, with the stem cut open to reveal the wispy central cord and there's a closer view here ![]() . In a fresh stem the cord is more like a smooth string, without those loose, wispy strands. The cord is present in the other species that the DNA evidence groups with Coprinus comatus - but the cord is absent from those Coprinus species that are not grouped with Coprinus comatus. This cord within the Coprinus comatus stem is not a recent discovery, for it had been shown clearly in a painting of Coprinus comatus, published in 1781 in a book by the French naturalist Jean Baptiste Francois' Pierre' Bulliard (1752-1793), but the phenomenon of inkiness was taken to be more significant than the presence of that cord.
. In a fresh stem the cord is more like a smooth string, without those loose, wispy strands. The cord is present in the other species that the DNA evidence groups with Coprinus comatus - but the cord is absent from those Coprinus species that are not grouped with Coprinus comatus. This cord within the Coprinus comatus stem is not a recent discovery, for it had been shown clearly in a painting of Coprinus comatus, published in 1781 in a book by the French naturalist Jean Baptiste Francois' Pierre' Bulliard (1752-1793), but the phenomenon of inkiness was taken to be more significant than the presence of that cord.
The upshot of all this is that there is still a genus Coprinus, but it has a much narrower definition than in the past and contains just a few species. Well over 90% of the old Coprinus species have been placed in the genera Coprinellus, Coprinopsis or Parasola. Of these Coprinellus and Coprinopsis are resurrections in that these generic names had been in use during the late 1800s for some species that a little later were subsumed in the genus Coprinus. The genus Parasola was established in 2001 to accommodate various 'Coprinus' species that did not fit the new concept of Coprinus and could not be accommodated in Coprinellus, Coprinopsis nor in any other existing genera. Incidentally, those responsible for establishing the genus Parasola were Scott Redhead, Rytas Vilgalys and John Hopple so, given earlier comments about the ICN requiring authors' names to be associated with taxa, this taxon name would be cited in full as Parasola Redhead, Vilgalys & Hopple .
.
Classification and identification - final words
While classification must bring together many different strands of evidence (using a variety of methods), for identification it is most sensible to use whatever features make it easiest to answer the question: What's the name of the specimen in front of me? Earlier on this page I said that a feature that is of little value in classification can still be very useful in identification. The converse also holds in that evidence important in a classificatory scheme may be of little practical use in the mundane task of specimen identification. At first glance these may seem strange statements but the important fact is that, while ideas of how fungi are classified may change, which will sometimes involve a change of name, the fungi themselves don't change. Inkiness no longer means the specimen is a species of Coprinus, but inkiness is still such an uncommon phenomenon that it remains a very useful identification tool, but one that needs to be augmented. While inkiness would no longer take you to just one genus, it would take you to a small group of genera, after which you'd use additional evidence to determine the genus. One piece of additional evidence you might use is the presence or absence of a hyphal cord, a feature more practical for identification purposes (since it is easily seen with the naked eye) than the DNA sequences that led to the changes in classification.
On the MACROSCOPIC FEATURES page you'll find a variety of features that are both useful in identification and easy to see.
![An Australian Government Initiative [logo]](/images/austgovt_brown_90px.gif)